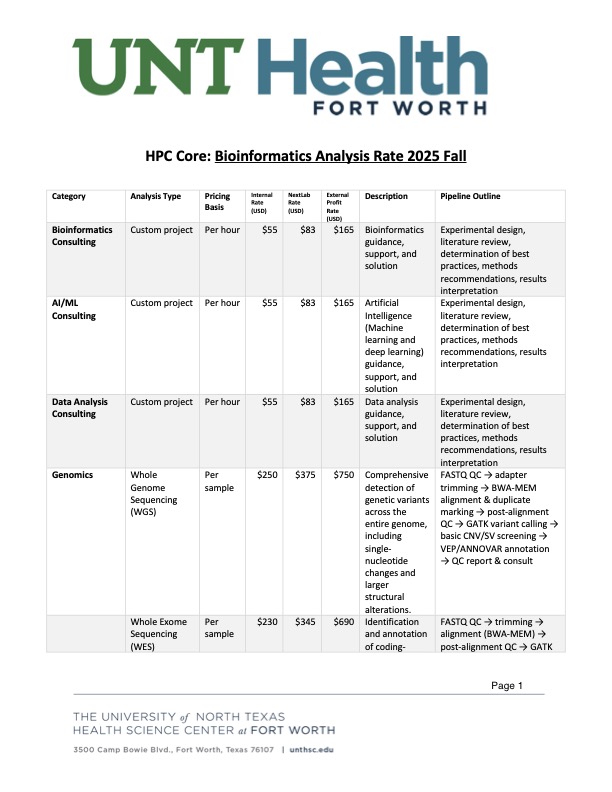
 Data Science & Computing (DSC) Core
Data Science & Computing (DSC) Core
The Data Science & Computing (DSC) Core at UNT Health aims to deliver computational
support for research applications, including big-data analytics, bioinformatics, statistical
support and AI/ML-driven research development.
The DSC Core is a high-throughput computing resource to support a wide array of biomedical
research across multiple disciplines. Some of the complex analyses, including but
not limited to are genomics, transcriptomics, proteomics and metabolomic studies,
as well as structural biology, drug discovery and systems biology. Applications range
from whole genome sequencing and single-cell transcriptomics to molecular modeling,
molecular docking and artificial intelligence-driven research. Additionally, the Core
supports clinical and translational research by facilitating clinical trial data processing,
Bayesian modeling, adaptive trial designs, survival analysis and other advanced computational
methodologies to foster innovation across the entire UNT Health research enterprise.
The system setup consists of compute nodes featuring a multi-node architecture with
AMD EPYC 9754 processors, each equipped with 256+ CPU cores, supporting parallel processing
for computationally intensive workflows. The GPU acceleration will utilize NVIDIA
H100 Tensor GPUs, optimized for deep learning, molecular simulations and AI-driven
analytics. The DSC Core will also include cold storage with a 1 PB infrastructure,
designed for long-term archival and management of large datasets. With state-of-the-art
computational resources, cloud integration options and scalable infrastructure, the
DSC Core is committed to enabling data-driven discoveries across diverse scientific
disciplines at UNT Health.
iLab
Log in to iLab to see a full list of equipment and schedule time in the DSC Core.



 Data Science & Computing (DSC) Core
Data Science & Computing (DSC) Core